Translate this page into:
Clinicopathological features and BRAFV600E mutation analysis in colorectal cancer
* Corresponding author: Dr. Usha Pai K.N, Department of Pathology, Jubilee Mission Medical College and Research Institute, Thrissur, Kerala, India. ushadev@gmail.com
-
Received: ,
Accepted: ,
How to cite this article: Thomas N, Pai K.N U, Raju L, Sureshkumar R, Ragitha T. S. Clinicopathological features and BRAFV600E mutation analysis in colorectal cancer. Asian J Oncol. 2025;11:6. doi: 10.25259/ASJO_60_2024
Abstract
Objectives
To assess the clinicopathological parameters in colorectal cancer. To determine the proportion of BRAFV600E mutations among colorectal cancer (CRC) patients.
Material and Methods
The cross-sectional study was conducted in 85 consecutive samples of histologically confirmed colon/rectal adenocarcinoma cases from a Tertiary Care Hospital, Thrissur, for eighteen months. Informed consent was obtained from patients whose specimens were collected, and the tests were performed after approval from the Institutional Ethics Committee. All the gross findings were recorded, including the size of the specimen, the tumor site, and the gross appearance. Histological sections were studied under a light microscope. Further deoxyribonucleic acid (DNA) isolation and amplification refractory mutation system- polymerase chain reaction (ARMS PCR) were carried out to confirm the BRAF mutation status. Data were entered into MS Excel and analyzed using IBM-SPSS software version 25.
Results
55.3% (47 cases) of CRC were above 65 years, 63.5% (54 cases) were males and 36.5% were females (M: F 3:1). Majority of tumors were located in the sigmoid colon (31.8%) followed by rectum (16.5%), ascending colon (14.1%), hepatic flexure (10.6%), caecum (7.1%), transverse colon (5.9%), descending colon (4.7%), and splenic flexure (1.2%). 91.8% of cases were low grade, and 8.2% were high grade. 50.6% of tumors were pT3, and the least were T4 (2.4%). Out of 85 study participants, BRAF was positive in 2(2.4%) patients and negative in 83(97.6%) patients, respectively.
Conclusion
This is the first such South Indian study. The small sample size, demographic disparities in the study population, and pre-analytical factors could explain the low positivity number. In the future, routine molecular testing for various biomarkers including BRAF may become inevitable in targeted therapy of CRC for practicing precision medicine.
Keywords
ARMS PCR
BRAF V600E mutation
Colorectal cancer
South Indian study
INTRODUCTION
Colorectal cancer (CRC) is the third most common cancer worldwide, causing considerable cancer-related deaths in both sexes.[1] The prevalence of colorectal cancer varies globally due to factors such as demographics, ethnicity, sedentary lifestyle, culture, genetics, and intake of high carbohydrate and low fiber diets. Over the last few years, the global incidence of CRC is about 151030; affecting 53.4% of men and 46.57% of women.[2] There is variation in the incidence rate of CRC within Asia, being uniformly low in all South Asian countries and high in all developed Asian countries.[3] In India, the incidence rates of CRC in males and females are 4.3 and 3.4 per 100,000, respectively, but recent studies have shown an increasing trend in the Indian population.[4]
CRC is a divergent disease resulting from multifarious genomic and epigenomic aberrations involving several signaling pathways. Based on the genetic mutation, CRC can be categorized as inherited, familial, and sporadic type, of which sporadic cases form a major share.[5] Three main molecular pathways due to several mechanisms are identified in the pathogenesis of CRC. 1. Chromosomal instability (CIN). 2.Microsatellite instability (MSI). 3. CpG island methylator phenotype (CIMP) pathway.[6,7] The BRAF (Rat sarcoma virus RAF and B-Raf proto-oncogene) gene is a valuable biomarker among the various genetic mutations.[8] BRAF-mutated CRC is associated with the worst overall (OS) survival and is considered a negative prognostic marker of CRC compared to wild-type BRAF.[9] According to the English literature on CRC among Asian countries, India has the highest BRAF prevalence rate of 11.1%, and the lowest number of 1.1% is reported from Taiwan. In contrast, China has a heterogeneous distribution of genetic mutations.[10] As per the location, the colon has the highest rate of BRAF mutations at 67.9% and a higher incidence of 59.9% in an advanced stage of cancer.[10] The BRAF oncogene is found on chromosome arm 7q34, which has 18 exons and codes for a 766 amino acid peptide with a molecular weight of 84 kDa. It produces a serine/ threonine kinase, a vital part of the RAS/RAF/MEK/ERK signaling cascade that encourages cell growth.[11,12] Numerous mutations have been identified in BRAF, the most prevalent of which is the Thymine to Adenine transversion (c.1799T>A) inside codon 600, which results in the amino acid level substitution of valine by glutamate (p.V600E). BRAFV600E mutation is seen in nearly one in ten patients with advanced colorectal cancer and approximately accounts for 5 to 9 % of the total cases. Some studies have reported that the V600E substitution is linked to specific clinicopathological characteristics such as proximal localization, microsatellite instability, mucinous histology, and chemotherapy resistance.[7,13,14] Non-V600E BRAF mutations are seen in younger CRC patients with a better prognosis.[15] In India, a literature survey has shown few reports from North India and no reports from South India.[15-19] In the present study, we analyzed the prevalence of BRAF mutations in CRC and correlated them with clinicopathological characteristics.
MATERIAL AND METHODS
The present study was conducted in the Department of Pathology, in a Tertiary Care Institution in South India, for eighteen months after approval from the Institutional Ethics Committee (IEC Ref No: 72/19/IEC/JMMC&RI). Informed consent was obtained from all the patients included in the study whose specimens were received in the department, and the Declaration of Helsinki procedures were followed. Formalin-fixed paraffin-embedded (FFPE) samples (85) of primary CRC were selected. All other non-epithelial malignant tumors of the colon and patients who received neoadjuvant radiotherapy/chemotherapy were excluded.
Histopathological examination
The resected specimens were fixed in 10% formol saline overnight. All the details, such as the procedure performed, dimensions of the specimen, site, size, and appearance of the tumor, margins, and lymph nodes were recorded. Tissue was processed in a semi-automated tissue processor followed by paraffin embedding. The 5-micron sections were stained using a standard Hematoxylin and Eosin staining protocol. All the sections were studied under a light microscope for the histological type and grade. Tumors were staged according to the eighth edition of the American Joint Committee on Cancer (AJCC) – tumor, node, metastasis (TNM) classification system.
Mutation analysis
Genomic DNA was extracted from 8 sections at 10µm thickness. DNA was extracted from the formalin fixed paraffin (FFP) using the Kit (Qiagen, Hilden, Germany) method and stored at -20°C. ARMS-PCR amplification was carried out by using the Primer pairs; forward outer (FO)-5’-CTCTTCATAATGCTTGCTCTGATAG-3’, reverse outer (Ro) - 5’-GCCTCAATTCTTACCATCCAC-3’, forward wild (FW) -5’-GTGATTTTGGTCTAGCTACAGT-3’, reverse mutant (RM) -5’-CCCACTCCAATCGAGATTTCT-3’.[20] The polymerase chain reaction (PCR) reaction was carried out with standardized conditions such as; initial denaturation at 95°C for 5 min, followed by 40 cycles of denaturation at 94°C for 30 sec, annealing at 59°C for 20 sec, and extension at 72°C for 20 sec with a final extension at 72°C for 5 minutes. The PCR product was then checked in 2% agarose gel.
Statistical analysis
Data were entered into MS Excel and analyzed using IBM-SPSS software version 25. P-value <0.05 was considered statistically significant. Numerical variables were expressed as Mean and Standard deviation. Clinicopathological parameters were expressed as Frequency and Percentages. A Chi-square test was used to compare the relationship between the grade and size of the tumor.
RESULTS
Characteristics of study subjects
Of the total 85 cases, 54 (63.1%) were men and 31 (36.5%) were women. The average age distribution of patients was 9 (10.6%) less than 45 years, 29 (34.1%) between 45 to 65 years, and 47 (55.3%) were more than 65 years of age.
Histopathological evaluation and molecular analysis
The size distribution of tumors was 0-5 cm in 45 (52.9%), 6-10 cm in 36 (42.4%), and 11-15cm in 4 (4.7%) cases. In our study majority of tumors were located in the sigmoid colon (31.8%), followed by the rectum (16.5%), ascending colon (14.1%), hepatic flexure (10.6%), caecum (7.1%), transverse colon (5.9%), descending colon (4.7%), and splenic flexure (1.2%). Regarding the tumor grade, 78 (91.8%) were low-grade tumors, and 7 (8.2%) were high-grade. The p-value for tumor grade and size was not statistically significant (P=0.638). In this study, pT3-43 (50.6%) was the most frequent tumor pT stage among the 85 study participants, while pT1- 2 (2.4%) was scarce. pN0- 43 (50.6%) was the most common nodal status, while pN2- 2 (2.4%) was infrequent. BRAF mutation analysis was performed in all cases. In our study, only 2.4% (2 patients) were positive for the BRAFV600E mutation, and the remaining were negative. [Figures 1 and 2]
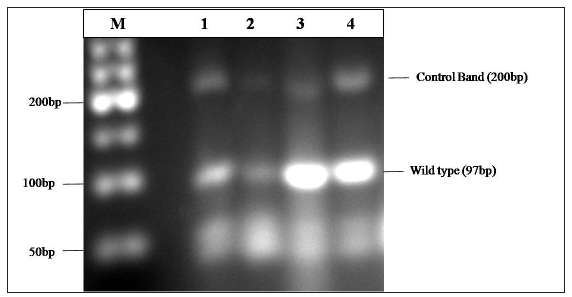
- Representative ethidium bromide-stained agarose gel showing wild type (97bp) and control (200bp) bands. Marker (M) is of 50bp ladder mixed with 2X gel loading buffer. Patient samples: Lanes1,2,3 and 4.
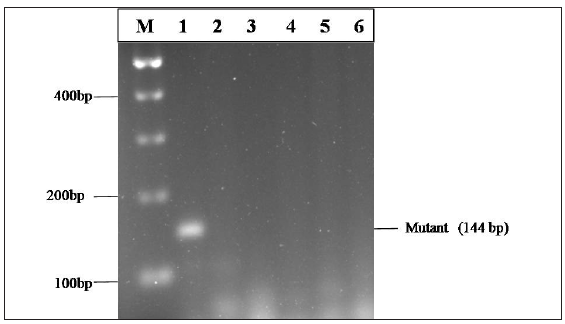
- Representative ethidium bromide-stained agarose gel showing BRAF mutation (144bp). Marker (M) is of 100bp ladder mixed with 2X gel loading buffer. Lane 1: Positive case. Lanes 2 – 6: Negative cases.
DISCUSSION
CRC is the third most common cancer leading to cancer-related mortality globally. Early diagnosis of colorectal cancers, surgical treatment, and response to adjuvant therapy have contributed to better outcomes in patients. The revelation of different signaling pathways in CRC has revolutionized the therapeutic perspective due to the use of combined targeted therapy, thereby prolonging the overall survival period.[21] Mainly, three diverse molecular pathways: the chromosomal instability pathway (mutations in P53 and KRAS), the mutator pathway (loss of function of DNA mismatch repair proteins), and the serrated pathway (BRAFV600E mutation) lead to CRC. Among these, the BRAFV600E mutation is usually observed in CRC with microsatellite instability (MSI). Screening is essential as patients harboring the BRAF mutation have an aggressive outcome and are unlikely to have Lynch syndrome.[22]
In our study, patients older than 65 years (55.3%) were the predominant age group involved. Gender-wise, CRC was greater in males (63.1%) than in females (36.5%), which is identical to the earlier reports from India and worldwide.[10,22-24] In this study, the age and gender of the patients did not show any statistically significant correlation. This disparity could be attributed to the varying sample sizes employed in different individual studies. The difference in male-to-female incidence in CRC is pertinent in prognostication, as males have a poorer outcome.[10] As the genetic pathways of CRC in the proximal and distal colon are divergent while some mutations are mutually exclusive, the location of the tumors is crucial.[25] In our cohort, the sigmoid colon (31.8%) was the preponderant site, followed by the rectum (16.5%), which is comparable to previous reports from Asia, including India.[22] There are isolated Indian reports of a higher number of right-sided colonic cancer and rectal cancer in younger individuals.[26,27] According to the ethnicity of the study samples, the worldwide BRAF mutation rate ranges from 7- 10%.[8]
In this study, BRAFV600E mutation analysis was performed on all 85 cases, of which two cases were positive (2.4%).[16] This is the same as the Indian study report by Lashkar et al on rectal cancers.[15] Other Indian reports showed varying numbers of BRAF positivity; the highest number of 21% of BRAF mutation in CRC is reported by R. Eachkoti et al.[17-19,23] The Asian population shows a lower frequency of BRAFV600E mutation compared to the Caucasian population.[28] However, one report from Iran recorded a complete lack of BRAF mutation in CRC.[29] Our results of low BRAF positivity concur with overall low incidence in Asia compared to the West. In this study, we found BRAF mutation in each male and female patient with low-grade tumors located in the right and left colon, respectively, in contrast to the previous report of BRAF mutation commonly found in females >70 years and the right colon.[30] Patil et al.[31] found a correlation of BRAF with low-grade carcinomas.[31] As T tumor stage (BRAFV600E mutation frequency is substantially greater in stage II/III than in stage IV) is still debatable in terms of how well it predicts outcomes in early disease, and more research is needed.[30] The majority (50.6%) of the cases in our study had pT3 tumors, and the least cases (2.4%) had T4 tumors, which should alert the treating physician to BRAF mutational testing. A survey by Lie et al.[32] reported a strong association of BRAF mutation with clinicopathological parameters like age, gender, location, histological type, and TNM staging, which was conspicuously absent in our study similar to the previous Indian data.[23]
The role of BRAF mutation in the prognosis of CRC is well recognized and is typically poorer.[12] Due to the increasing significance of this mutation, the National Comprehensive Cancer Network Guidelines now recommend BRAF mutation testing in patients with metastatic disease.[33] Moreover, BRAF mutant metastatic CRCs present a unique metastatic pattern, showing high rates of peritoneal metastases and distant lymph node metastases.[34] In the present study, there was no metastasis recorded in both BRAF-positive cases. Additionally, BRAF mutant metastatic CRC patients frequently show an early resistance to targeted therapy, and only about half of these patients respond to second-line chemotherapy, suggesting that more aggressive and individualized combined therapies may be effective in selected patients’ cohorts.[34] Recent studies have proved the conjunctive efficacy of targeted therapy using EGFR and BRAF inhibitors in metastatic CRC.[35] There is only one documented Indian report of Non-BRAF V600E mutation.[17]
On follow-up with our BRAF-positive patients, the first female patient, an elderly person, did not receive chemotherapy and succumbed to colonic cancer during the pandemic, five months after the right hemicolectomy. The second male patient on colostomy following sigmoid resection was deferred chemotherapy due to comorbidities, such as uncontrolled diabetes mellitus, hypothyroidism, cirrhosis liver, and renal failure. Four years later, he died following liver metastasis.
The limitation of this study was the small sample size. A collaborative study of molecular tests in a large Indian population is necessary for explicit mutational data in CRC.
CONCLUSION
This is the first South Indian cohort study of CRC on BRAF mutation. The small sample size, demographic disparities in the study population, and pre-analytical factors could explain the low positivity number. In the future, routine molecular testing for various biomarkers, including BRAF may become inevitable in targeted therapy of CRC for practicing precision medicine.
Author contributions
N.K.T: Involved in study design, experiments and drafting the manuscript, U.P.K.N: Participated in conceptualization, study design, and overall supervision, L.R and S.K.R: Experimental trouble shooting and manuscript editing, R.T.S: Experimental trouble shooting and manuscript drafting.
Ethical approval
The research/study approved by the Institutional Review Board at Institutional Ethics Committee Jubilee Mission Medical College & Research Institute, number 72/19/IEC/JMMC&RI, dated 14th November 2019.
Declaration of patient consent
The authors certify that they have obtained all appropriate patient consent.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
Use of artificial intelligence (AI)-assisted technology for manuscript preparation
The authors confirm that there was no use of artificial intelligence (AI)-assisted technology for assisting in the writing or editing of the manuscript and no images were manipulated using AI.
REFERENCES
- Statin uses and mortality in colorectal cancer patients: An updated systematic review and meta-analysis. Cancer Med. 2019;8:3305-13.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893-917.
- [CrossRef] [PubMed] [Google Scholar]
- A genetic model for colorectal tumorigenesis. Cell. 1990;61:759-67.
- [CrossRef] [PubMed] [Google Scholar]
- Microsatellite instability in colorectal cancer. Br J Surg. 2006;93:395-406.
- [CrossRef] [PubMed] [Google Scholar]
- Colorectal cancer in India: controversies, enigmas, and primary prevention. Indian J Gastroenterol. 2011;30:3-6.
- [CrossRef] [PubMed] [Google Scholar]
- Targeting RAS-ERK signaling in cancer: promises and challenges. Nat Rev Drug Discov. 2014;13:928-42.
- [CrossRef] [PubMed] [Google Scholar]
- Molecular mechanisms of colon cancer progression and metastasis: Recent insights and advancements. Int J Mol Sci. 2020;22:130.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Clinicopathologic features and prognosis of BRAF mutated colorectal cancer patients. Front Oncol. 2020;10:563407.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- A systematic review and meta-analysis on the occurrence of biomarker mutation in colorectal cancer among the asian population. Biomed Res Int. 2022;2022:5824183.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Molecular pathology of colorectal cancer. Adv Anat Pathol. 2020;27:20-6.
- [CrossRef] [PubMed] [Google Scholar]
- The molecular pathogenesis of colorectal cancer and its potential application to colorectal cancer screening. Dig Dis Sci. 2015;60:762-72.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Mutations of the BRAF gene in human cancer. Nature. 2002;417:949-54.
- [CrossRef] [PubMed] [Google Scholar]
- BRAF-mutated colorectal cancer: Clinical and molecular insights. Int J Mol Sci. 2019;20:5369.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Rectal cancer profiling identifies distinct subtypes in India based on age at onset, and genetic, epigenetic, and clinicopathological characteristics. Mol Carcinog. 2015;54:1786-95.
- [CrossRef] [PubMed] [Google Scholar]
- The prevalence of BRAF, PIK3CA, and RAS mutations in indian patients with colorectal cancer. South Asian J Cancer. 2022;11:190-4.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Prevalence and coexistence of KRAS, BRAF, PIK3CA, NRAS, TP53, and APC mutations in indian colorectal cancer patients: Next-generation sequencing-based cohort study. Tumour Biol. 2017;39:1010428317692265.
- [CrossRef] [PubMed] [Google Scholar]
- A study of BRAF mutation in colorectal carcinoma in indian population. J Cancer Res Ther. 2018;14:1403-6.
- [CrossRef] [PubMed] [Google Scholar]
- Prevalence and prognostic relevance of brafV600E mutation in colorectal carcinomas from kashmir (North India) valley. Mutagenesis. 2018;33:225-30.
- [CrossRef] [PubMed] [Google Scholar]
- Sensitive detection of BRAF V600E mutation by amplification refractory mutation system (ARMS)-PCR. Biomark Res. 2013;1:3.
- [CrossRef] [PubMed] [Google Scholar]
- Novel targeting approaches and signaling pathways of colorectal cancer: An insight. World J Gastroenterol. 2018;24:4428-35.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Histopathological study of neoplastic lesions of large intestine in Kashmir Valley, India. Int J Res Med Sci. 2014;2:1097.
- [CrossRef] [Google Scholar]
- Molecular spectrum of KRAS, BRAF, and PIK3CA gene mutation: Determination of frequency, distribution pattern in indian colorectal carcinoma. Med Oncol. 2014;31:124.
- [CrossRef] [PubMed] [Google Scholar]
- Emerging trends in colorectal cancer: Dysregulated signaling pathways (Review) Int J Mol Med. 2021;47:14.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- The chromosomal instability pathway in colon cancer. Gastroenterology. 2010;138:2059-72.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]
- Rectal cancer in young adults: a series of 102 patients at a tertiary care center in India. Colorectal Dis. 2009;11:475-9.
- [CrossRef] [PubMed] [Google Scholar]
- Rectal cancer in young indians--are these cancers different compared to their older counterparts? Indian J Gastroenterol. 2014;33:146-50.
- [CrossRef] [PubMed] [Google Scholar]
- An ecological study of trends in cancer incidence and dietary changes in hong kong. Nutr Cancer. 1997;28:289-301.
- [CrossRef] [PubMed] [Google Scholar]
- BRAF gene mutation analysis in colorectal cancer in the south of Iran. Iranian J Colorectal Res. 2014;2
- [Google Scholar]
- Objective criteria for the grading of venous invasion in colorectal cancer. Am J Surg Pathol. 2010;34:454-62.
- [CrossRef] [PubMed] [Google Scholar]
- Clinico pathological correlation with mutation profiling of colorectal cancer for KRAS, BRAF,NRAS and PIK3CA genes in Indian patient cohort. Ann Oncol. 2015;26 Suppl 9:42-70.
- [Google Scholar]
- Limited evidence of non-response bias despite modest response rate in a nationwide survey of long-term cancer survivors—results from the NOR-CAYACS study. Journal of Cancer Survivorship. 2019;13:353-63.
- [CrossRef] [PubMed] [Google Scholar]
- Optimizing targeted therapeutic development: analysis of a colorectal cancer patient population with the BRAF(V600E) mutation. Int J Cancer.. 2011;128:2075-84.
- [PubMed] [Google Scholar]
- Lymph node recoveries from 2427 pT3 colorectal resection specimens spanning 45 years: Recommendations for a minimum number of recovered lymph nodes based on predictive probabilities. Am J Surg Pathol. 2002;26:179-89.
- [CrossRef] [PubMed] [Google Scholar]
- EGFR-mediated re-activation of MAPK signaling contributes to the insensitivity of BRAF mutant colorectal cancers to RAF inhibition with vemurafenib. Cancer Discov. 2012;2:227-35.
- [CrossRef] [PubMed] [PubMed Central] [Google Scholar]







